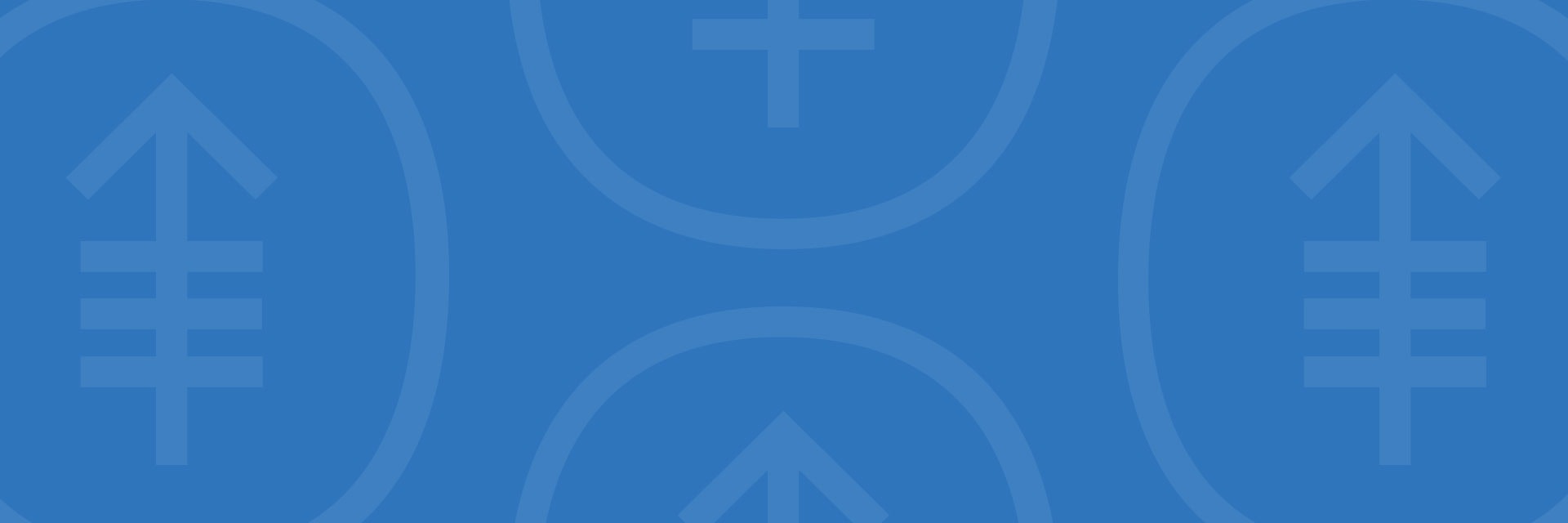
The Minkui Luo Lab
Research

The Luo Lab is interested in developing and leveraging novel approaches at the interface of chemistry and biology to annotate functional roles of epigenetic modulators. Epigenetic regulation is essential for maintaining cellular fates and its dysregulation has been associated with various diseases including cancer. Among key epigenetic modulators are diverse protein methyltransferases/demethylases and methyl-arginine/lysine (Rme/Kme) effector proteins. With novel chemical tools developed in-house, we aim to systematically interrogate Rme/Kme-involved epigenetics with emphasis on novel molecular mechanisms and biological outcomes. The Luo laboratory is also interested in designing and synthesizing epigenetic inhibitors with the emphasis of the distinct modes of action in disease-relevant contexts. Researchers in the Luo Lab with expertise in chemistry, biochemistry and biology work in a highly collaborative manner to develop unprecedent chemical reagents for biological discovery, elucidate disease-causing mechanisms with molecular details, and explore novel treatments against cancer.

Featured News

Publications Highlights
Cai XC, Zhang T, Kim EJ, Jiang M, Wang K, Wang J, Chen S, Zhang N, Wu H, Li F, Dela Seña CC, Zeng H, Vivcharuk V, Niu X, Zheng W, Lee JP, Chen Y, Barsyte D, Szewczyk M, Hajian T, Ibáñez G, Dong A, Dombrovski L, Zhang Z, Deng H, Min J, Arrowsmith CH, Mazutis L, Shi L, Vedadi M, Brown PJ, Xiang J, Qin LX, Xu W, Luo M. A chemical probe of CARM1 alters epigenetic plasticity against breast cancer cell invasion. Elife. 2019 Oct 28;8:e47110. This work was highlighted by Elife Digest.
Chen S, Kapilashrami K, Senevirathne C, Wang Z, Wang J, Linscott JA, Luo M. Substrate-Differentiated Transition States of SET7/9-Catalyzed Lysine Methylation. J Am Chem Soc. 2019 May 22;141(20):8064-8067.
Chen S, Wiewiora RP, Meng F, Babault N, Ma A, Yu W, Qian K, Hu H, Zou H, Wang J, Fan S, Blum G, Pittella-Silva F, Beauchamp KA, Tempel W, Jiang H, Chen K, Skene RJ, Zheng YG, Brown PJ, Jin J, Luo C, Chodera JD, Luo M. The dynamic conformational landscape of the protein methyltransferase SETD8. Elife. 2019 May 13;8:e45403. This work was highlighted by Elife Digest, News of Shanghai Institute of Materia Medica, MSKCC Blog, and TPCB Webpage.
People

Minkui Luo, PhD
- Chemical Biologist Minkui Luo develops cutting-edge tools, technologies and concepts to annotate functions of protein-posttranslational modifications and designs inhibitors for cancer therapies.
- BS, Fudan University
- PhD, Princeton University
- [email protected]
- Email Address
- 646-888-3066
- Office Phone
- Download CV
- PDF File
Members









































Achievements
- Maximizing Investigators’ Research Award (R35), NIGMS/NIH (2019)
- Eli Lilly Award in Biological Chemistry, American Chemical Society (2015)
- Clinical & Translational Science Center Novel Award, Weill Cornell Medical College (2014)
- Basil O'Connor Starter Scholar, March of Dimes Birth Defects Foundation (2011)
- Director’s New Innovator Award, National Institutes of Health (2010)
- Alfred W. Bressler Scholar, Alfred W. Bressler Scholars Endowment Fund (2010)
- The V Scholar Award, V Foundation for Cancer Research (2009)
- Outstanding Postdoctoral Research Prize, Albert Einstein College of Medicine (2007)
- Best Presentation Award, 3rd ACS Annual Metropolitan New York Area Poster Program for Graduate Students in the Chemical Sciences (2003)
- Rohm & Haas Fellowship, Fudan University (1998)
- Du Pont Fellowship, Fudan University (1998)
Open Positions
To learn more about available postdoctoral opportunities, please visit our Career Center
To learn more about compensation and benefits for postdoctoral researchers at MSK, please visit Resources for Postdocs
Career Opportunities
Get in Touch
-
Lab Head Email
-
Office Phone
-
Lab Phone
Disclosures
Members of the MSK Community often work with pharmaceutical, device, biotechnology, and life sciences companies, and other organizations outside of MSK, to find safe and effective cancer treatments, to improve patient care, and to educate the health care community. These activities outside of MSK further our mission, provide productive collaborations, and promote the practical application of scientific discoveries.
MSK requires doctors, faculty members, and leaders to report (“disclose”) the relationships and financial interests they have with external entities. As a commitment to transparency with our community, we make that information available to the public. Not all disclosed interests and relationships present conflicts of interest. MSK reviews all disclosed interests and relationships to assess whether a conflict of interest exists and whether formal COI management is needed.
Minkui Luo discloses the following relationships and financial interests:
-
American Society for Biochemistry and Molecular Biology
Professional Services and Activities (Uncompensated) -
Central China Normal University
Professional Services and Activities -
Chinese-American Chemistry & Chemical Biology Professors Association (CAPA)
Professional Services and Activities (Uncompensated) -
Elsevier
Professional Services and Activities -
Federation of American Societies for Experimental Biology (FASEB)
Professional Services and Activities (Uncompensated) -
Gordon Research Conferences
Professional Services and Activities (Uncompensated) -
International Chemical Biology Society
Professional Services and Activities (Uncompensated) -
King Abdullah University of Science and Technology
Professional Services and Activities (Uncompensated)
-
Pacifichem, Inc.
Professional Services and Activities (Uncompensated) -
Peking University
Professional Services and Activities (Uncompensated) -
Research Grants Council (RGC), Hong Kong
Professional Services and Activities -
Shanghai Institute of Materia Medica, Chinese Academy of Sciences
Professional Services and Activities (Uncompensated) -
Shenzhen Bay Laboratory
Professional Services and Activities -
The Mark Foundation for Cancer Research
Professional Services and Activities -
The University of Hong Kong
Professional Services and Activities -
Westlake University
Professional Services and Activities
The information published here is a complement to other publicly reported data and is for a specific annual disclosure period. There may be differences between information on this and other public sites as a result of different reporting periods and/or the various ways relationships and financial interests are categorized by organizations that publish such data.
This page and data include information for a specific MSK annual disclosure period (January 1, 2024 through disclosure submission in spring 2025). This data reflects interests that may or may not still exist. This data is updated annually.
Learn more about MSK’s COI policies here. For questions regarding MSK’s COI-related policies and procedures, email MSK’s Compliance Office at [email protected].
