Transmembrane sensing of host immune effectors: The Rip1 and Chalkophore pathways
Signal transduction across membranes through regulated intramembrane proteolysis is conserved throughout all domains of life. In the SREBP pathway of human cells, the membrane-bound SREBP transcription factors undergo sequential cleavage by site-one and site-two proteases (S1P and S2P, respectively), with human S2P being the founding member of the zinc metalloprotease family of intramembrane proteases. RseP, an E. coli S2P, cleaves the anti-Sigma factor for SigE and thereby also regulates transcription in response to environmental signals, specifically unfolded outer membrane proteins. Our original characterization of the Rip1 (Rv2869c) pathway in M. tuberculosis revealed that this S2P is a critical virulence determinant.
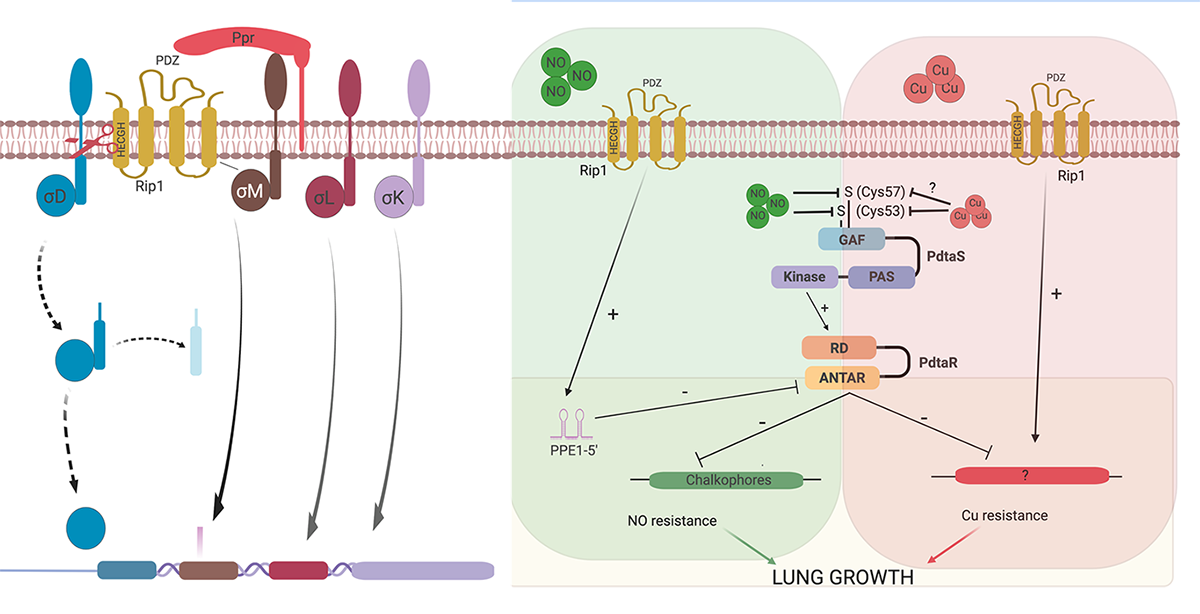
We have identified four anti-Sigma factor substrates of Rip1, anti-SigK, L, M, and D which regulate multiple downstream target genes. However, the virulence functions of Rip1 are independent of these sigma factor pathways. Recent work (Buglino et al, Elife 2021) has identified a new pathway downstream of Rip1 controlled by the PdtaS/PdtaR two component system that controls NO and Cu resistance through a complex set of interacting positive and negative feedback loops. Ongoing work in the lab seeks to understand the molecular details of this signal transduction system that is critical for Mtb virulence.
We have recently focused on the diisonitrile chalkophore system of Mtb. Diisonitrile lipopeptides are produced by Mtb and bind Copper ions with high affinity. Our work has shown that this system allows Mtb to resist copper deprivation both in culture and in the host. This chalkophore acquired copper maintains the copper centers of the Mtb heme:Cu respiratory oxidases, defining a new mechanism by which bacterial pathogens resists host nutritional immunity.
Studies in human TB
In collaboration with our colleagues at Weill Cornell Medicine and the GHESKIO Center in Port Au Prince, Haiti, and with the support of the NIAID’s Tuberculosis Research Unit Program, we are studying the effect of antimycobacterial antibiotics on intestinal microbiome composition and the relationship of those microbiome changes to TB disease resolution.



