What are the functions of 3′UTRs?
To identify genes whose functions depend on their 3′UTRs, we are deleting 3′UTR sequences from endogenous gene loci and investigate the resulting phenotypic changes. We study the biological functions of 3′UTRs mostly during iPS cell differentiation into various cell types.

What is the mechanism by which 3′UTRs control protein activity independently of protein abundance?
We found that 3′UTRs can regulate protein activity of transcription factors and chromatin regulators in a manner that is independent of protein abundance. 3′UTR-dependent activity regulation occurs during translation of mRNAs in mRNA-based cytoplasmic compartments. We are studying mechanistically how mRNAs act as chaperones for proteins during translation and how mRNAs change protein conformations.
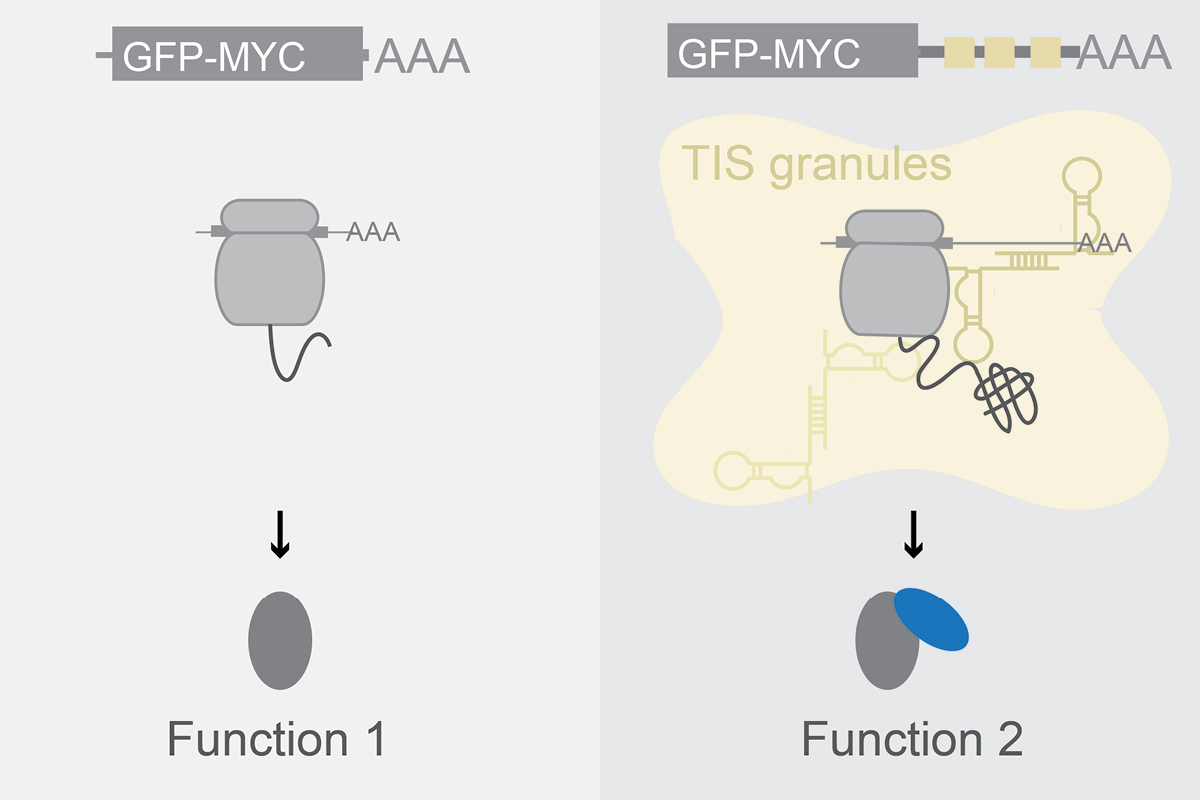
What are the structural and regulatory roles of mRNAs in the cytoplasm?
We found that the cytoplasm is compartmentalized into the cytosol and several mRNA-based translation environments. These translation environments are generate by specific RNA-binding proteins, such as TIS11B or FXR1 (see figure), together with their bound mRNAs and can be considered large mRNP networks. As the underlying scaffold of these networks is composed of mRNAs, our findings indicate that mRNAs fulfil structural roles in the cytoplasm. Going forward, we want to identify the assembly principles of TIS granules, the FXR1 network, and a currently poorly characterized translation environment generated by TIAL1 on the ER surface.
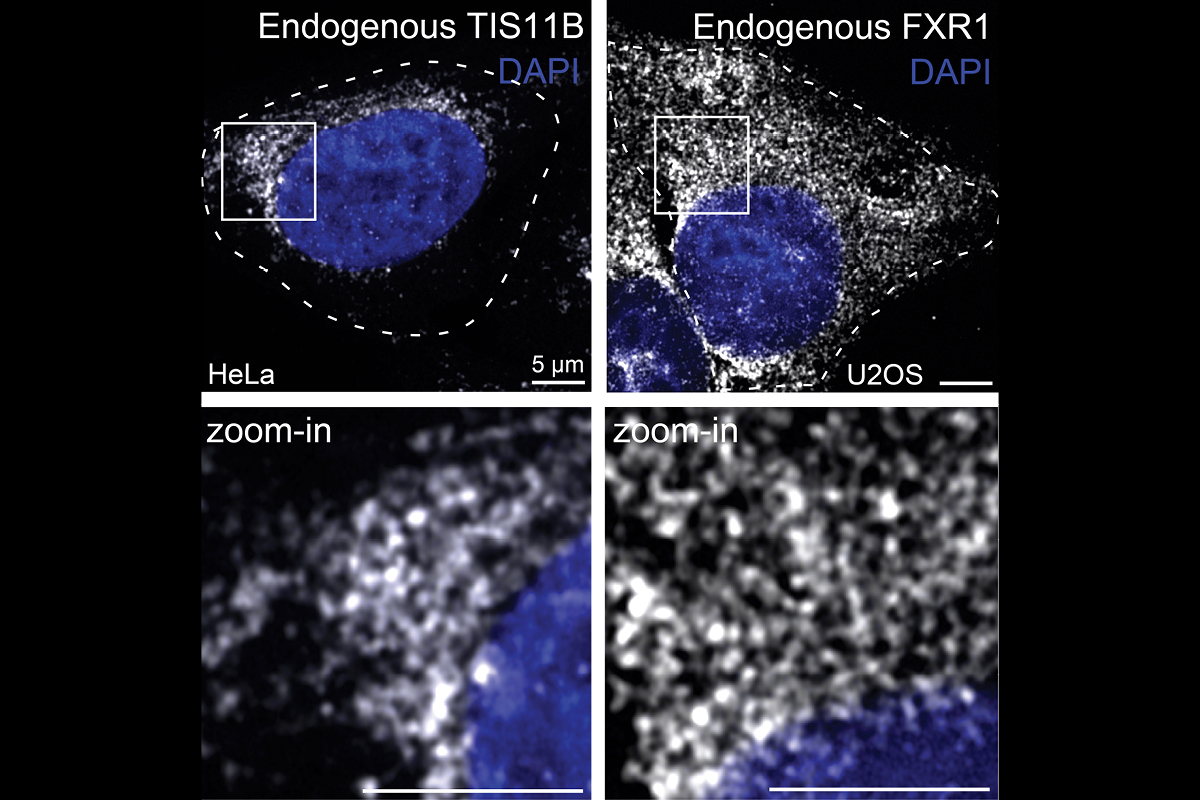
What happens in mRNA-based translation environments, such as TIS granules or the FXR1 network?
We found that mRNA-dependent protein complexes are generated when mRNAs are translated in TIS granules. We want to learn what determines the specificity of mRNA-dependent protein complexes. Are these complexes mostly determined by the surrounding mRNAs or by the enriched proteins in the translation environments?
Can we use our insights to design novel RNA therapeutics?
We propose that we can use nucleic acid-based small molecules to activate or inhibit protein functions. Moreover, these small molecules might even be able to target separate protein functions.