The Massagué lab investigates metastasis stem cells and their stromal niches throughout the metastatic cascade, with a growing interest in the dormant phase of metastasis. We are defining the phenotypic plasticity and evolution of metastatic cell populations from dormancy to outbreak, identifying drug targets, and enabling clinical trials to treat metastasis. We entered this area on realizing the central role of TGF-β and its interplay with oncogenic signals in metastasis. We collaborate with biologists, data scientists, and clinicians to leverage these findings for clinical benefit.
TGF-β signaling in development and disease
The TGF-β regulatory system plays crucial roles in the preservation of organismal integrity by simultaneously regulating different cell types and cellular functions. TGF-β signaling controls embryo development, tissue homeostasis, and injury repair through coordinated effects on cell proliferation, phenotypic plasticity, migration, metabolic adaptation, and immune surveillance of multiple cell types in shared ecosystems. Defects of TGF-β signaling disrupt immune tolerance, promote inflammation, underlie the pathogenesis of fibrosis and cancer, and contribute to the resistance of these diseases to treatment. Having elucidated the TGF-β signaling pathway, we are investigating how cells interpret TGF-β signals depending on the presence of context-dependent SMAD cofactors and chromatin configurations. Recent work uncovered a major interface between the TGF-β and RAS-MAPK pathways. TGF-β and RAS, signaling through SMAD and RAS-responsive element-binding protein 1 (RREB1) respectively, jointly activate a multi-arm regulatory program in epithelial progenitors and carcinoma cells (Su et al Nature 2020; Lee et al Cell 2024). RREB1 localizes to acetylated histone H4 marks in histone H2A.Z-loaded nucleosomes in cell plasticity genes, priming these enhancers for activation by a TGF-β activated SMAD4-INO80 nucleosome remodeling complex. These findings illuminate the operation of a bifunctional program that promotes metastatic outgrowth.
Phenotypic regulation
Epithelial-to-mesenchymal transitions (EMTs) are phenotypic plasticity processes that confer migratory and invasive properties to epithelial cells during development, wound-healing, and cancer. TGF-β is a potent inducer of EMTs implicated in liver disease, pulmonary fibrosis, and carcinoma metastasis. Lung adenocarcinoma (LUAD) cells undergo a TGF-β-dependent EMT associated with fibroblast activation and extracellular matrix remodeling. RAS-activated RREB1 primes enhancers of EMT and fibrogenic genes for activation by chromatin remodeling complexes that the TGF-β/SMAD pathway recruits to these enhancers. Both the EMT arm and the fibrogenic arm of this response program are essential for pulmonary metastasis in LUAD models. Inhibiting RREB1 disables this pro-metastatic process.
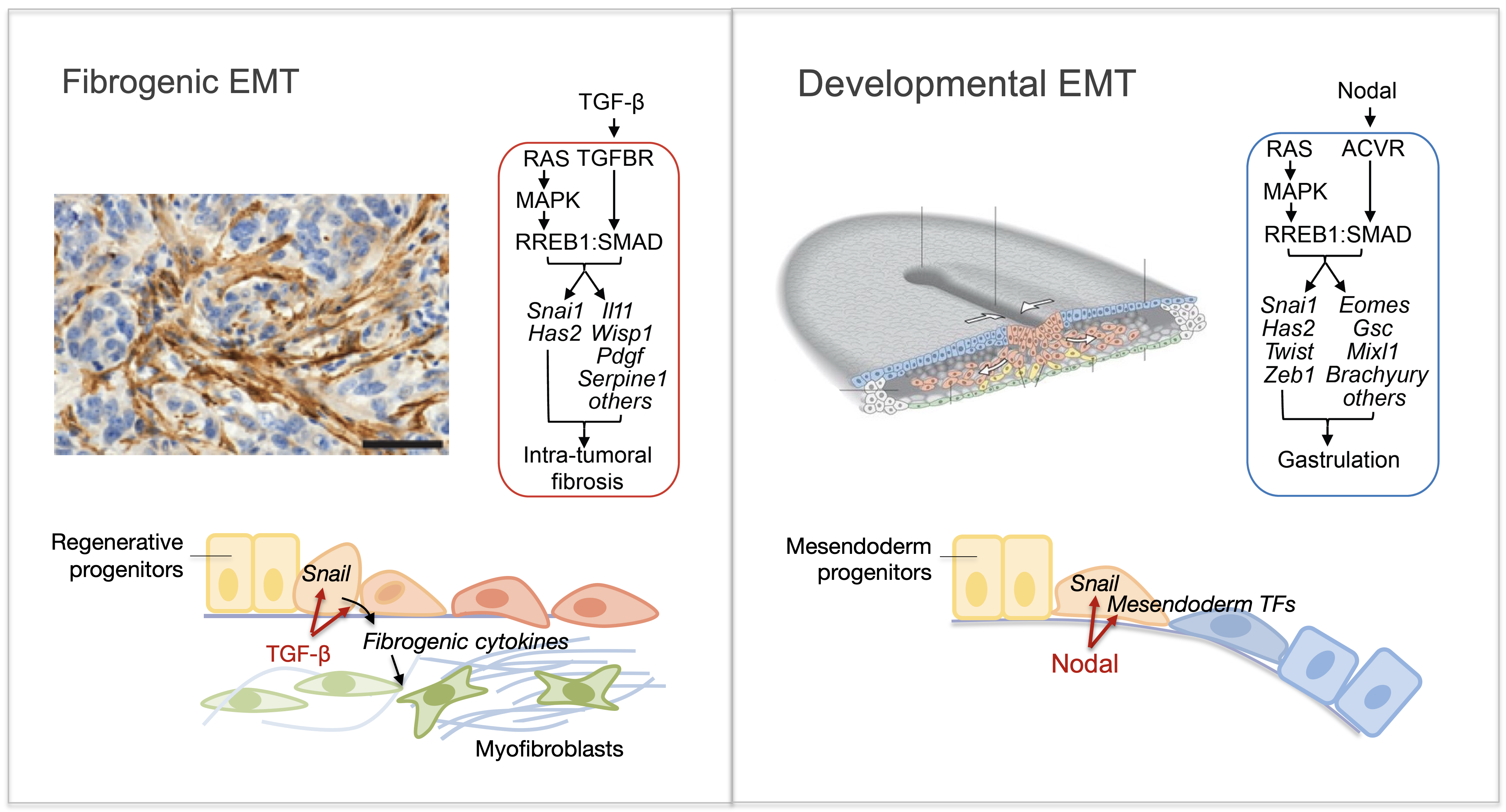
In primitive, stem-like SOX2+ LUAD progenitors, TGF-β induces growth arrest accompanied by a full EMT response that subsequently transitions into an atypical mesenchymal state of round morphology and lacking actin stress fibers. TGF-β drives this long-term transition by inducing the expression of gelsolin, which converts a stress fiber-rich mesenchymal phenotype into a cortical actin-rich spheroidal state of low biomechanical stiffness to protect metastatic stem cells from killing by CD8 T cells and NK cells. Thus, LUAD primitive progenitors undergo an atypical EMT as part of a strategy to evade immune-mediated elimination (Wang et al Nature Cancer 2026). We are building on these insights to gain a better understanding of epithelial plasticity regulation by TGF-β in development, fibrosis and metastasis.
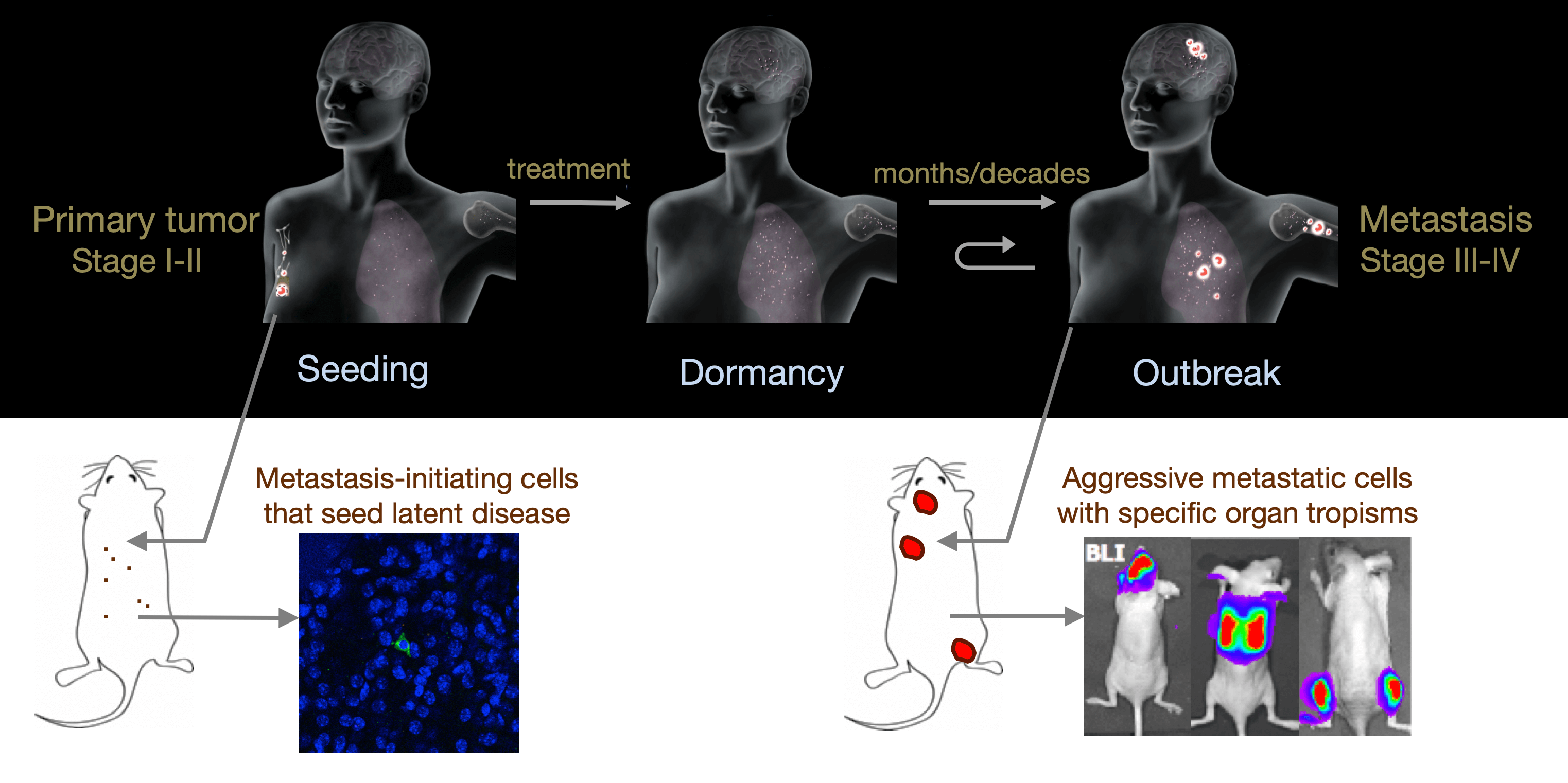
From metastatic dormancy to outbreak
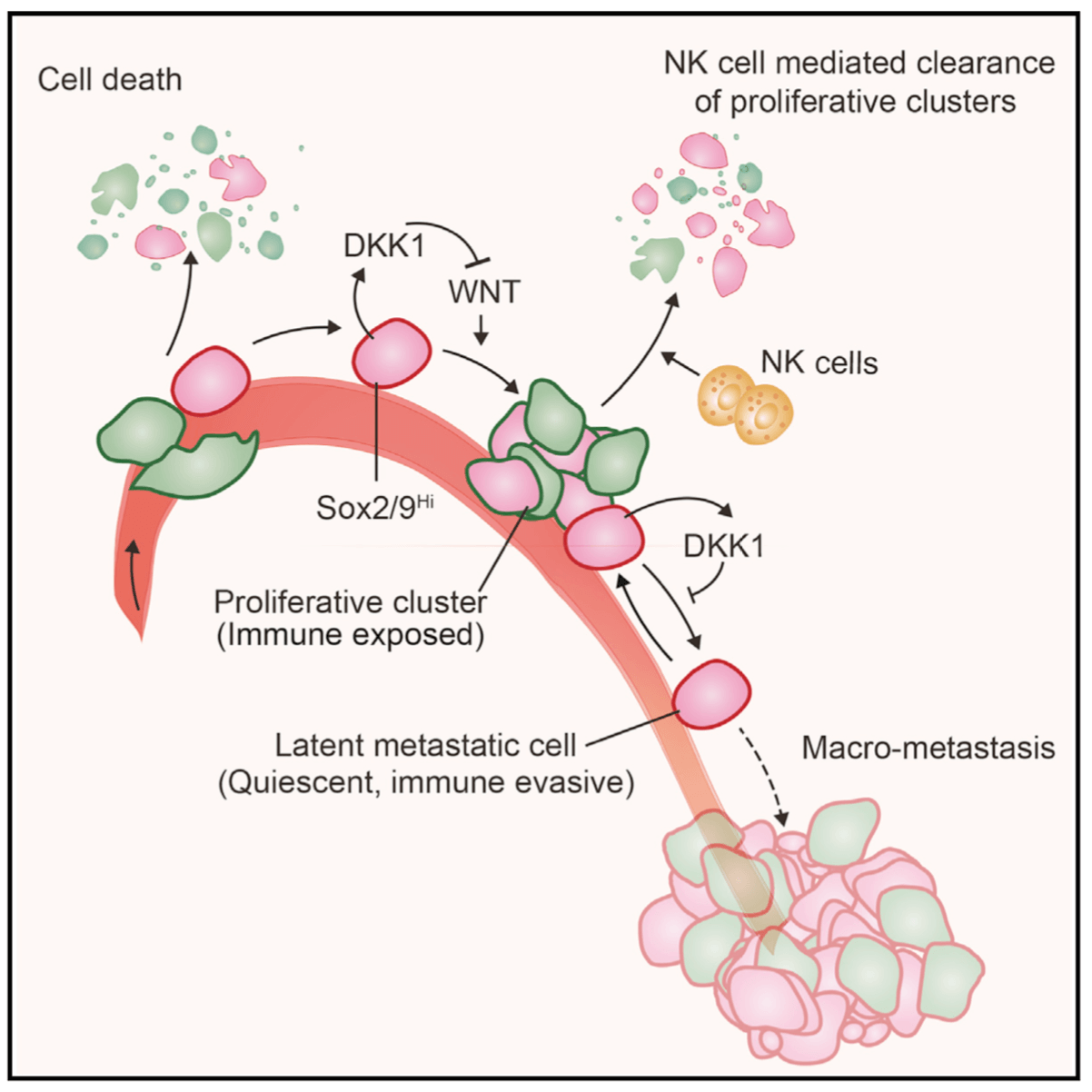
Metastasis is a complex, multiorgan, and often fatal process (accounting for 90% of cancer-related deaths). Cancer cells that disseminate from a tumor to distant sites enter a period of dormancy that may persist from months to decades before giving rise to detectable metastasis. Adjuvant therapy treatments seek to prevent overt metastasis by eliminating residual malignant cells during this dormancy period. Efforts to improve adjuvant therapy are hindered by an insufficient understanding of the molecular mechanisms that preserve the long-term viability of dormant metastatic cells. Identifying these mechanisms is needed to improve treatments and prevent relapse. Using mouse models that the lab developed, we are investigating the evolution of SOX2+ metastasis stem cells that enter a dormant phase after infiltrating target organs, remain viable under immune surveillance, and eventually reinitiate tumor growth. This work revealed roles of TGF-β as a driver of immune evasive metastatic dormancy and reveal vulnerabilities of metastatic cells. We uncovered key roles of STING (Hu et al Nature 2023) and L1CAM in this process (Park et al bioRxiv 2025) that can be exploited for the eradication of residual disease.

Organ-specific metastasis
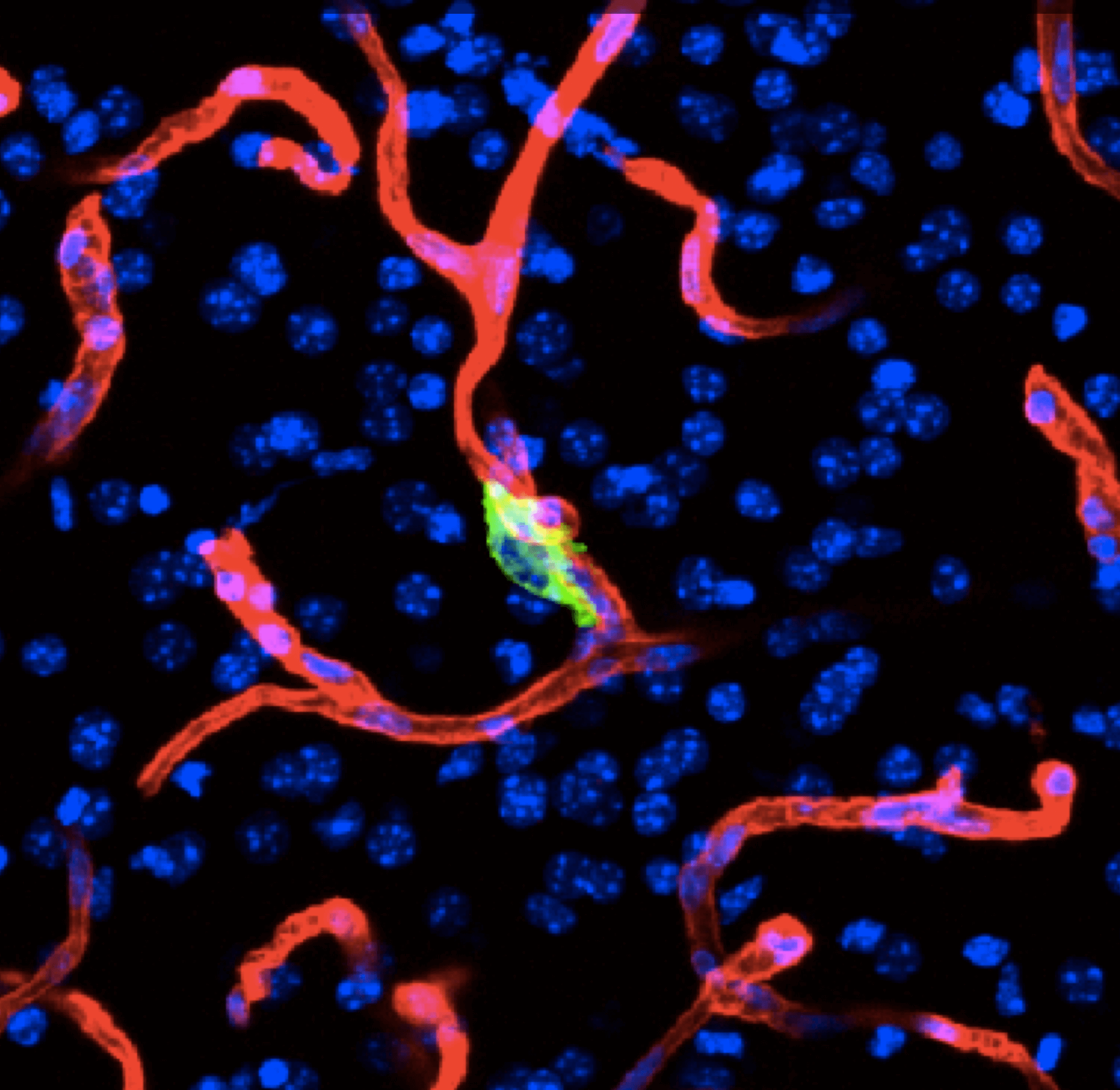
During metastasis, disseminated cancer cells undergo profound, tissue-specific reprogramming that reshapes their phenotype, metabolism, and therapeutic responses, giving rise to organ adaptations and organ tropisms characteristic of each type of cancer. The lab has identified a series of mediators of organ-specific metastasis and is currently focusing on the specific case of brain metastasis. Brain metastasis is highly lethal. Its incidence is ten-fold higher than that of all other brain tumor types combined. We created mouse models of brain metastasis from lung cancer and breast cancer and are using these models to identify relevant mediators of brain metastasis. In recent work we demonstrated that distinct tumor architectures and microenvironments for the initiation of metastasis in the brain (Gan et al Cancer Cell 2024). This work argues for a precision oncology framework that integrates these organ site-specific programs into treatment design.