Epigenetics — the study of biochemical changes that influence a cell’s genetic programming — is arguably one of the fastest-moving fields of cancer research. At Memorial Sloan Kettering, scientists are exploring a number of new drugs that target epigenetic processes. If such drugs prove to be safe and effective in patient trials, they might eventually offer a groundbreaking treatment tactic.
Unlike most existing cancer drugs, which work by destroying cancer cells, some epigenetics-based therapies have the potential to reorient the diseased cells and set them on a path back to normal growth and development. We recently reported on one of these therapies, AG-221, which is being investigated in an early-stage clinical trial in patients with leukemia and other blood disorders. Now the promise of another experimental leukemia drug called a DOT1 inhibitor is on the rise as well.
Until recently, our researchers believed the DOT1 inhibitor might be useful in only one aggressive subtype of acute myelogenous leukemia (AML), representing about 15 percent of AML cases. But a recent study shows the drug might be effective against additional AML subtypes that together account for about half of the close to 19,000 new cases of the disease diagnosed each year in the United States.
The research also sheds light on epigenetic changes in blood cells that may lead to AML— and this knowledge could translate into more-powerful therapies. To learn more we talked to physician-scientist Scott Armstrong, who led the study, published in the journal Cancer Cell late last year, and is the head of MSK’s newly launched Center for Epigenetics Research.
What is epigenetics?
Epigenetics literally means “beyond DNA.” The term refers to the fact that all the cells in our bodies — brain cells, liver cells, and so on — have the exact same DNA, but each cell type uses its DNA differently by turning on different sets of genes at different times.
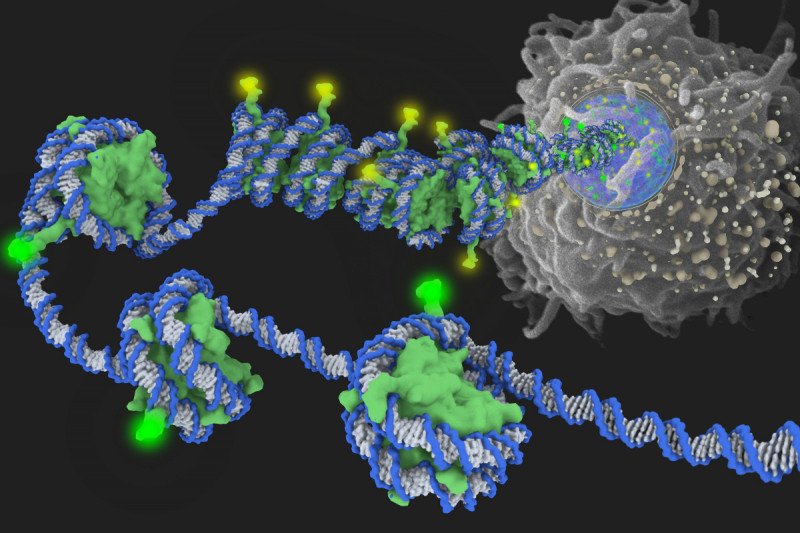
This programming is determined by the way a cell’s DNA is spatially organized. Inside the nucleus of a cell, DNA strands are wrapped around spool-shaped aggregates of proteins called histones. This packaging is extremely dense; if a cell’s DNA became unwrapped and stretched out it would measure six feet!
Broadly speaking, there are two types of epigenetic changes:
- Biochemical changes of histone proteins either tighten or relax the wrapping of DNA. When a stretch of DNA is bound tightly around histones, genes located within that region do not turn on.
- DNA methylation, a change on the DNA strand, can also affect how genes are expressed.
For each of the body’s cell types, epigenetic modifications manifest across the genome in a unique and dynamic pattern. We might think of this pattern as software that orchestrates the activity of thousands of genes.
Why is epigenetics relevant to cancer?
When changes take place in the proteins that control epigenetic processes, this can affect the expression of numerous genes that play a role in cancer initiation, development, or progression. Cancer-promoting genes that are normally silent may become activated while other genes that can suppress cancer are shut off.
The field of cancer epigenetics really took off in the last couple of years, once we learned that one of the most commonly mutated processes in cancer is the one by which histones and DNA are chemically modified. The discovery caused enormous excitement among researchers because histone and DNA modifications can potentially be reversed with drugs.
We know of about 70 different ways histones can be modified. By better understanding how the processes work, we might be able to tap into untold opportunities to develop drugs that restore normal gene expression in cancer cells.
Your recent study suggests using a drug to inhibit DOT1 might be effective against many different types of AML. How does this drug work?
In patients with a subtype called MLL-rearranged AML, immature blood cells fail to develop properly because a group of genes called HOX genes — which normally are shut off — become active, causing the cells to proliferate uncontrollably. DOT1, the target of the drug, is an enzyme that alters certain chemical groups on a histone protein.
Earlier work in my lab has shown that the DOT1 inhibitor can be used to turn off HOX gene expression in MLL-rearranged AML cells. And by shutting off the HOX genes, we can thwart the cells’ cancerous behavior and make them grow up into mature, functional blood cells.
Now, in our recent paper, we show that the drug could potentially help many more patients than we initially thought, which is very exciting. This is because a number of AML subtypes besides MLL-rearranged AML are driven by HOX gene expression.

Are there other potential implications of the study?
Yes. We were able to clarify the inner workings of the epigenetic switch that turns on HOX genes in leukemia cells. We found that this switch can be manipulated by modifying a single carbon atom.
This is pretty remarkable. Just imagine that the biological process that leads to a person’s leukemia might be heavily influenced by a tiny shift of one atom!
We’re now working to use this information to understand more about the control of HOX gene expression and design more-effective drugs against cancers that depend on these genes. We are lucky to be collaborating with MSK structural biologist Dinshaw Patel and his lab members, who have made a number of groundbreaking discoveries in the field of epigenetics.