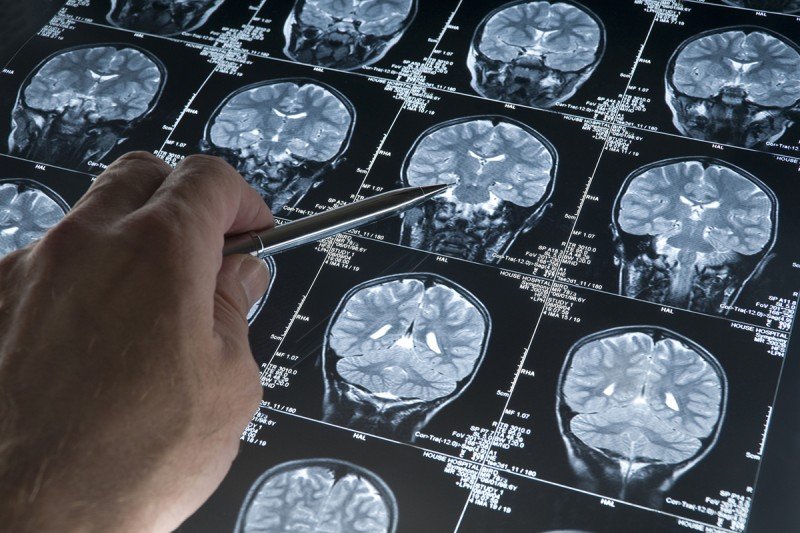
There are more than 100 subtypes of brain tumors, making them a challenge to accurately diagnose. This week, an international team led by researchers at the German Cancer Research Center is announcing they have developed a new way to classify these tumors. A paper describing the tool was published in Nature.
Proper diagnosis and classification is vital for brain tumors. It not only helps with prognosis but also enables doctors to determine the best treatment. For example, some tumor types respond better to radiation therapy than others, while some respond to certain chemotherapy drugs. Some don’t need to be treated at all and can just be closely monitored.
We spoke with Memorial Sloan Kettering Pediatric Neuro-Oncology Service Chief Matthias Karajannis, who participated in the research, about why this study is important and what it means for people with brain cancer, both at MSK and around the world.
Why are brain tumors so hard to classify?
Brain tumors, especially in children, are not a uniform disease. They are actually many different diseases. Until recently, these tumors were diagnosed exclusively by looking at cells under a microscope. But even for the most experienced pathologist, it can be challenging to tell some of the different types apart. You could have three pathologists look at the same tumor sample and come up with three different diagnoses.
Advances in gene sequencing, such as MSK-IMPACTTM, have led to improvements. But there are limitations with sequencing cancer-related mutations, especially in pediatric brain tumors. Many of the subtypes don’t have a known gene mutation that is commonly linked with them that can be used to classify the subtype of tumor.
What’s different about the system that’s been developed now?
This research team has developed a completely new classification system. It is the first time anyone has shown a way to reliably distinguish from among the 100-plus different types of brain tumors.
The system looks at what is called the tumor’s methylation profile. Once the profile is determined in the lab, it can be fed into an algorithm in a computer and automatically matched with samples that already exist in the database. This approach is based on the fact that each tumor subtype has a different methylation profile.
Another aspect that was important about the study is that we demonstrated this approach is equally feasible and reliable across multiple cancer centers.
What is a “methylation profile”? How is it different from other types of tumor analysis?
Methylation is a way that DNA is modified without changing the sequence of the four DNA letters. It’s one of the factors that influences how and when genes are translated into proteins.
I like to use a music analogy. The DNA sequence is the notes on the page. The methylation profile helps determine how fast or slow the music is played, how loud or how soft.
When I came to MSK, I was pleased to learn that the Molecular Diagnostics Service, under the leadership of Service Chief Marc Ladanyi, was already establishing the technical platform to study the methylation profile of tumors. The equipment needed to do these kinds of studies had just arrived. Dr. Ladanyi and his team will be working with the New York State Department of Health to help methylation profiling become an approved clinical test, the first step toward insurance coverage. It’s currently considered an experimental test for research purposes.
Will this study affect how MSK diagnoses its patients?
We are already using this tool with all of our pediatric brain tumor patients, and with many adults who have brain tumors as well. We’ve been using it for a while. But now we’re in the process of working out an agreement with the German Cancer Research Center team so that we’ll be able to do all the analysis with the tool right here at MSK in Molecular Diagnostics. Currently, we send them the results from our methylation studies and they send back the information about tumor classification.
We are also looking at how methylation profiling can be used to diagnose other types of tumors, especially sarcomas. Sarcomas are another kind of tumor that have many, many different subtypes. And just like in brain tumors, these subtypes can be hard to distinguish based on how they look under the microscope, or even by looking at their genetic profile.
You came to MSK just over a year ago. Why did you decide to join the team?
I completed my fellowship training at MSK and then spent nine years at New York University School of Medicine. It was an exciting opportunity to return as Chief of the Pediatric Neuro-Oncology Service and to work with the world-renowned experts here, many of whom were part of my training.
MSK has one of the largest, if not the largest, pediatric neuro-oncology programs in the country. We see about 100 children who have been newly diagnosed with brain and spine tumors every year.
In addition, the research capabilities here are outstanding. Combining the infrastructure and equipment that was already here with my own work in molecular diagnostics made my coming here a perfect storm of opportunity. We are able to diagnose and treat people with brain cancer better than at any time before, and MSK is doing it better than any other institution.
What made you want to become a pediatric neuro-oncologist?
I went to medical school at a time when rapid progress was being made for treating other pediatric cancers, especially leukemia. There also were many new tools for diagnosing leukemia. By the time I graduated, the majority of children with blood cancers were being cured, thanks to better and more refined chemotherapy treatments.
I didn’t see the same progress being made for brain tumors. Techniques in surgery and radiation were getting better, and there was some limited success with chemotherapy. But there was still a lot of work to be done. Because I had lab training in molecular pathology, I knew that in the future, improving diagnosis would be a real game-changer in this field. And it has been. It’s been exciting and rewarding to be part of this quantum shift.