CUT&RUN
a novel chromatin profiling strategy as an alternative to ChIP. In CUT&RUN, unfixed permeabilized cells are incubated with target-specific antibody, followed by binding of a protein A/G-Micrococcal Nuclease (pAG/MNase) fusion protein to release specific protein–DNA complexes into the supernatant for paired-end sequencing. Compared with ChIP, CUT&RUN requires fewer cells and fewer reads, has a higher signal-to-noise ratio and higher resolution, has no solubility and fragmentation bias, and is faster.
Reference: Skene, P.J., J.G. Henikoff, and S. Henikoff, Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat Protoc, 2018. 13(5): p.1006-1019.
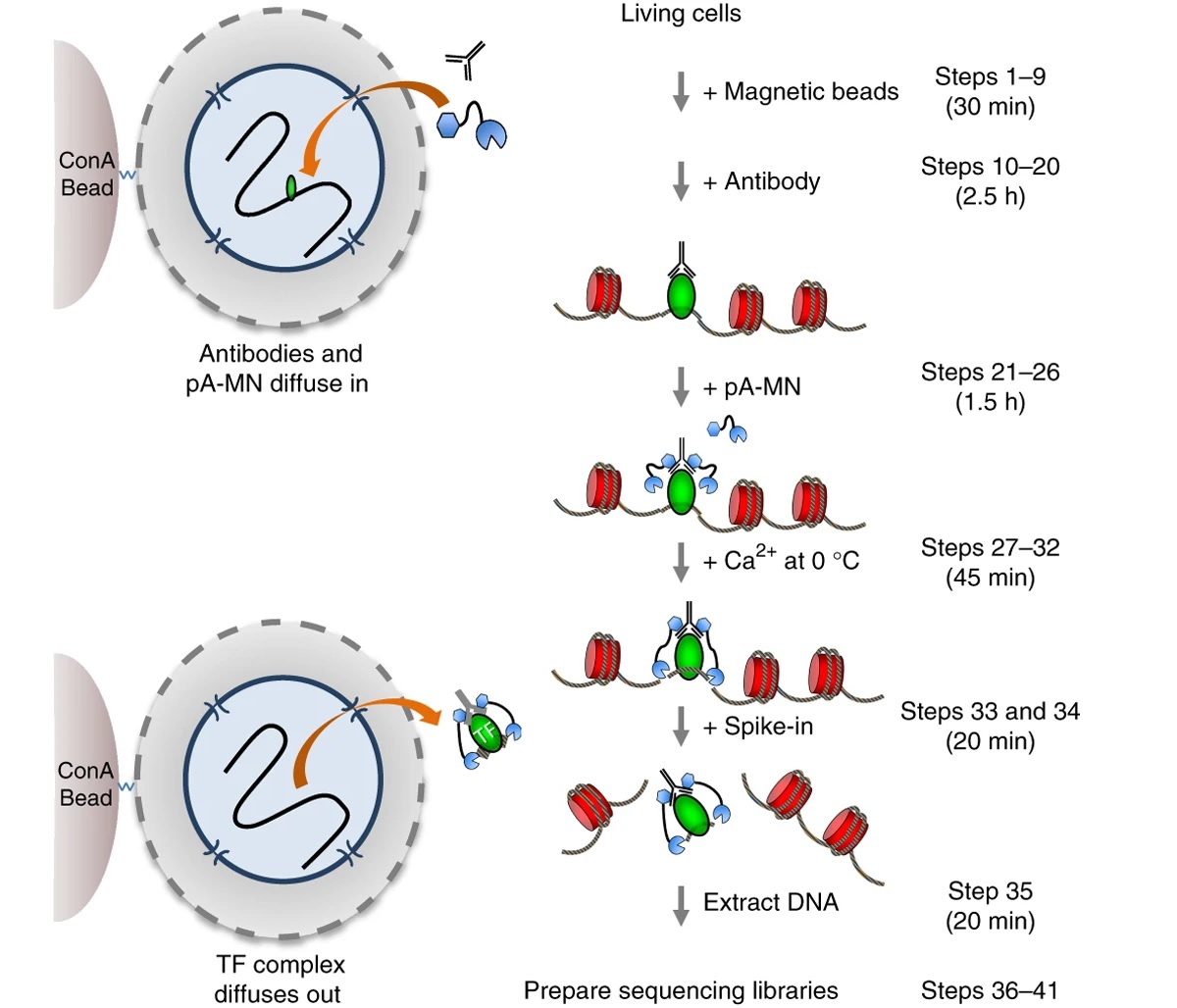
Image source