The Alan and Sandra Gerry Metastasis and Tumor Ecosystems Center is pleased to have funded the following MSK investigators and specific metastasis-related projects and resource acquisitions.

Ross Levine (Classic Individual)
Clonal hematopoiesis as a driver of metastatic relapse in early breast cancer
Efforts to identify genomic biomarkers to predict breast cancer outcomes have traditionally focused on tumor intrinsic factors; thus, identification of tumor extrinsic factors with prognostic and therapeutic importance is imperative to better predict patient outcomes and redefine therapeutic strategies for the prevention and treatment of metastatic breast cancer. Genomic sequencing studies have uncovered that 25% of breast cancer patients, carry somatic mutations in their leukocytes, a condition known as clonal hematopoiesis (CH). The incidence of CH increases with aging and has been recognized as a major risk factor for the development of subsequent hematologic malignancies and cardiovascular disease. CH mutations frequently occur in genes involved in myeloid malignancies, most commonly in the epigenetic modifier DNMT3A. A hallmark of breast cancer is the admixture of tumor cells and myeloid cells, which play a key role in determining therapeutic resistance and metastasis through the fine tuning of inflammatory signals. More recently, we have revealed that CH mutations are enriched in breast cancer tumor infiltrating leukocytes and that CH in putative cancer drivers (CH-PD) is associated with increased risk of solid tumor dependent morbidity and mortality. Although the most common cause of death in these patients was metastatic disease progression, the mechanisms by which CH contributes to therapeutic resistance and metastatic relapse remain unclear. Our central hypothesis is that CH alters the inflammatory responses of myeloid cells to drive therapeutic resistance and metastatic relapse via increased proliferation of disseminated cancer cells. This project will A) Determine why breast cancer cells re-emerge as metastasis by exploring the effects of CH-PD on the metastatic outcomes of patients with early breast cancer; and B) Determine how to prevent lethal metastatic recurrences by targeting inflammatory signals in mouse models of CH and breast cancer metastasis. This study represents an innovation because (1) the biological processes behind how aging and CH contribute to metastatic progression remains largely unexplored, and (2) understanding how CH mutant leukocytes contribute to the pro-metastatic process may provide a new tool to identify patients at highest risk of metastatic relapse and lead to the identification of actionable targets to limit tumor promoting inflammation and metastasis.

A. Ari Hakimi (Classic Individual)
Immunometabolic Coevolution as a Determinant of Response to Immunotherapy in ccRCC
Treatments combining inhibition of PD1 with inhibition of either VEGF or CTLA4 have revolutionized outcomes for patients with metastatic clear cell renal cell carcinoma (ccRCC), but only a fraction of patients respond to these therapies. Emerging evidence implicates the metabolism of both tumor and immune cells as critical mediators of sensitivity to combination immunotherapy via at least two axes. First, immunotherapy and anti-VEGF therapy directly alter the availability of key biosynthetic/energy-rich metabolites (glucose, glutamine) and signaling molecules (kynurenine, adenosine) delivered by the circulation to the microenvironment. Second, both immunotherapy and anti-VEGF therapy remodel the individual metabolic phenotypes of tumor and immune cells, including their expression of key metabolic transporters and catabolic enzymes and therefore their ability to compete for limited nutrients in the tumor milieu. These data raise the possibility that ccRCC evolution produces unique immunometabolic niches, defined jointly by specific patterns of metabolite availability and immune composition, that shape both initial sensitivity to immunotherapy and future molecular adaptation after exposure to immunotherapy. Our research group has pioneered the analysis of multimodal metabolomic and transcriptomic data in ccRCC to study the metabolic basis of tumorigenesis, metastasis, and therapeutic sensitivity. Based on this, we have (1) discovered novel metabolite phenotypes associated with existing, prognostically significant RNA signatures of the ccRCC TME (e.g. increased T-cell infiltration and NAD+ abundance) and (2) developed novel machine learning methods (tMIRTH) to impute metabolite abundance directly from RNA sequencing data. In parallel, we have (3) developed a novel immunocompetent mouse model of metastatic ccRCC that faithfully phenocopies ccRCC tumors and demonstrates robust responses to various immunotherapeutic regimens. The overarching hypothesis of this proposal is that the metabolism and immune microenvironment of ccRCC coevolve to produce unique microenvironments with defined sensitivities to immunotherapy. To address this hypothesis we will analyze bulk, multimodal metabolomic/microenvironmental profiling of ccRCC tumors (Aim 1) and the direct effect of immunotherapy on novel, immunocompetent, mouse models of ccRCC (Aim 2). Findings from our work will lead to the identification of novel metabolomic biomarkers for sensitivity to immunotherapy that can be measured prior to therapy and will reveal the full spectrum of metabolically-defined tumor microenvironments and illuminate the basic etiology of immunometabolic evolution in ccRCC and nominate rational metabolic interventions for combination strategies.

Morgan Huse (Classic Individual)
A Mechanoimmunological Basis for Metastatic Site Preference
Over the past decade, cytotoxic lymphocytes, comprising cytotoxic T lymphocytes (CTLs) and natural killer (NK) cells, have emerged as critical players in the control of metastatic outgrowth. Immunosurveillance by cytotoxic lymphocytes is generally conceived as a biochemical process in which tumor specific markers are recognized by activating receptors on the lymphocyte membrane. We have found, however, that immune cells also respond to the characteristic biophysical properties of cancer cells during metastasis. To colonize target organs, disseminated cancer cells activate a mechanotransduction pathway that enables them to spread and migrate along the abluminal surface of blood vessels. While this pathway is necessary for effective invasion of the metastatic niche, it also renders cancer cells more stimulatory to cytotoxic lymphocytes by rigidifying cell surface immunoreceptor ligands. Hence, the biophysical properties required for efficient colonization also sensitize metastatic cells to destruction by the immune system. Here, we will explore how this mechanical form of immunosurveillance, which we call mechanosurveillance, influences where metastatic growth occurs. Cellular mechanics are modulated continuously by cell-extrinsic biophysical signals. A particularly important manifestation of this crosstalk, called mechanoreciprocity, induces cells to adopt the mechanical properties of their immediate surroundings. Consistent with this principle, we and others have shown that cancer cells grown on stiffer substrates become stiffer themselves, whereas those grown on softer surfaces become softer. Whether environmentally-induced stiffening might sensitize cancer cells to mechanosurveillance in vivo, however, has not been explored. This an interesting question because metastatic microenvironments vary widely in their physical properties, ranging from very rigid (e.g. bone) to very soft (e.g. lung). Enhanced mechanosurveillance in rigid microenvironments would establish a regime in which cytotoxic lymphocytes control the spectrum of metastatic site preference by disproportionately suppressing outgrowth in organs like the bone. Using a murine model of metastasis, we have found that cancer cells colonizing the bone are significantly stiffer than cancer cells colonizing the lung, and that the in vivo expansion of bone metastasis is exquisitely sensitive to cytotoxic lymphocytes. Building on these preliminary observations, we propose that microenvironmental stiffness dictates the efficacy of mechanosurveillance and that this relationship shapes both metastatic site preference and the power of anti-tumor immunotherapy. We will investigate this hypothesis by applying materials science, functional assays, and single cell transcriptomics to mouse models of metastasis and patient samples. The successful completion of our Specific Aims could identify biomarkers for guiding antitumor immunotherapy and aid development of strategies for treating metastatic growth in specific target organs.

Scott Lowe (Classic Individual)
Somatic Deletions of type I IFNs in Immune Evasion and Metastasis
Deletions of chromosome 9p21.3 are pervasive across human cancers, and strongly associated with poor prognosis, altered immune infiltration, and resistance to immune checkpoint blockade. While the contributions of these deletions to cancer biology have been broadly ascribed to the disruption of the cell cycle inhibitors CDKN2A/B, approximately half of 9p21.3 deletions include a neighboring cluster of 16 type I interferon (IFN) genes. To determine whether IFN co-deletion contributes to the phenotypes associated with 9p21.3 loss, we leveraged a new genome engineering approach developed in our group termed Molecular Alteration of Chromosomes with Engineered Tandem Elements (MACHETE), which enables the rapid and flexiblegeneration of megabase-sized chromosomal deletions. By applying MACHETE to an immunocompetent mouse model of pancreatic ductal adenocarcinoma (PDAC), we found that tumors bearing concomitant loss of Cdkn2a/b and the IFN cluster exhibit reduced immune surveillance and enhanced metastatic spread due to evasion of CD8+ T cell surveillance. Motivated by these observations, this project aims to molecularly and functionally dissect the impact of IFN signaling and its disruption on the PDAC ecosystem in tumor progression and upon immunotherapy. We propose to refine our understanding of relevant activities in the 9p21.3 locus and to interrogate the immune environment and epithelial-immune cell interactions using latest-generation single cell methods. These studies will provide novel insights and broadly applicable knowledge on what may be the most frequent genetic mechanism of immune evasion in human cancer.

Karuna Ganesh (Classic Individual)
Molecular Mediators of Dynamic Cell State Plasticity in Colorectal Cancer Metastasis
Metastasis is the principal cause of cancer death. Recent studies have reveal that phenotypic plasticity, the dynamic adaptation of cancer cells to the stresses of dissemination and tumor regeneration in distant sites, is an overarching hallmark of metastasis. However, the molecular mechanisms that underpin the dynamic diversification and selection of cell states during tumor progression, and the ultimate endpoint cell states that are selected for during metastasis in advanced human solid tumors remain poorly understood. We have pioneered an integrated approach for single cell profiling and organoid derivation matched trios of synchronously resected normal colon, primary colorectal cancer (CRC), and CRC liver metastasis from patients undergoing cancer surgery at MSK. Our preliminary single cell analysis reveals distinct lineage trajectories that are selected for and against in the progression from normal epithelium to primary tumor and metastasis in the same patients. In this proposal, we will leverage our patient-derived organoid and orthotopic mouse models and cutting-edge single cell, live imaging, and computational approaches to dissect the transcription factors and lineage programs that underpin metastatic plasticity. Our work will illuminate fundamental mechanisms and clinically actionable signaling pathways underlying dynamic cell state transitions and lineage plasticity in metastasis, poised for clinical translation to improve cancer outcomes.
Simon Schwoerer (GMTEC Postdoctoral Researcher Innovation Award)
Investigating metabolism in the tumor stroma reveals an unexpected role of protein catabolism in pancreatic cancer
Pancreatic ductal adenocarcinoma (PDAC) is an aggressive tumor characterized by nutrient-deprivation in the tumor microenvironment. How PDAC cells continue to grow despite nutrient deprivation is not fully understood. In this project, we are going to investigate whether the of uptake and degradation of extracellular proteins in non-malignant stromal cells can support PDAC progression by proving an alternative source of free amino acids.
Francisco Sanchez-Rivera (GMTEC Postdoctoral Researcher Innovation Award)
Reconstructing complex cellular trajectories in vivo with single cell resolution
Reconstructing complex cellular trajectories in vivo with single cell resolution The overarching goal of this project was to develop and apply methods to 1) define the molecular trajectories that cells navigate in vivo during acquisition of their phenotypic identity, 2) investigate whether these trajectories exhibit significant spatiotemporal plasticity and dynamics, and 3) assess whether disease-associated genetic alterations (e.g. cancer mutations) influence these trajectories. While classical lineage tracing approaches have provided significant insight into these questions, achieving a complete and dynamic understanding of complex cellular trajectories and cell state transitions in vivo requires quantitative recording, measurement, and reconstruction of phylogenetic and transcriptional relationships over time with single cell resolution. Towards this goal, we developed an experimental framework in close collaboration with the group of Dana Pe’er that combines single cell RNA sequencing (scRNA-seq) and CRISPR-based genetic barcoding to dynamically characterize the emergence, plasticity, and lineage relationships of cancer cell populations in vivo. This system, which we refer to as Single-cell Transcriptomics And lineage Reconstruction (STAR), is based on a first-principles approach grounded on the hypothesis that each sgRNA has intrinsic editing properties largely determined by the sequence and difficult to predict without functional interrogation. This led us to develop and deploy a molecular recording ‘sensor’ platform (adapted from base editing sensor strategies partially supported by GMTEC; see Sánchez-Rivera et al., Nature Biotechnology, 2022) to identify sgRNAs with exceptionally high allele diversity and low large deletion probabilities that record at diverse editing rates. These optimized sgRNAs and cognate target sites are currently serving as a backbone to build an optimal lineage tracing system capable of achieving very high resolution and dynamic recording rates for robust lineage tracing over multiple temporal scales.
Ming Li (Classic Individual)
Reprogram Tumor-Associated Macrophages to Heal the Tumor Wound
Tumors develop by invoking a supportive microenvironment resembling a non-healing wound characterized by aberrant angiogenesis and chronic infiltration of tumor-associated macrophages (TAMs). Here we show that TAMs can be reprogrammed to heal the tumor wound following ablation of TSC1, a repressor of the metabolic regulator mTORC1. We found that the tumor parenchyma-localized TAMs exhibited low mTORC1 activity. The depletion of TSC1 in TAMs led to mTORC1-mediated tumor suppression independent of the adaptive immune system. Instead, TSC1-deficient TAMs redistributed to a perivascular niche and rectified the tortuous and hyperpermeable vasculature, causing tumor tissue hypoxia and cancer cell death in distant avascular regions. Notably, TSC1-deficient TAMs exhibited reduced expression of inflammatory and angiogenic genes but increased expression of transcripts that promote macrophage metabolism and scavenger function. Thus, TAMs can be metabolically reprogrammed via TSC-mTORC1 signaling to an anti-inflammatory and pro-resolving state to reconfigure the tumor vasculature and suppress cancer progression.
Minkui Luo (Classic Individual)
CARM1 Inhibition Alters the Epigenetic Plasticity of Metastatic Breast Cancer Cells
Numerous epigenetic events need to be orchestrated for breast cancer cells to metastasize to remote organs. Significant efforts have been made to identify the epigenetic cues that module metastatic outgrowth. CARM1 is an important epigenetic modulator to which metastatic breast cancer cells are addicted. High levels of CARM1 strongly correlate with breast cancer malignancy. We recently developed CARM1 chemical probes to interrogate CARM1-associated biology. The GMETC grant allowed us to evaluate in vivo efficacy of these compounds against breast cancer metastasis. While these treatments had no effect on the primary lung lesion, they significantly suppressed the secondary metastatic lesion and prolong overall survival. In addition, we conducted multiple sets of scRNA-seq analysis on gene expression profiles of MDA-MB-231 cells and develop in-house machine-learning algorithms dissect subpopulation-associated transcriptomic signatures of invasive cells. We concluded that the invasion capability of MDA-MB-231 cells mainly arises from an invasion-prone subset, 80% of which can be depleted by our CARM1 inhibitors. Differential expression analysis of scRNA-seq data further revealed the single-cell transcriptional signatures of metastasis-addicted genes such as MORC4, S100A2, RPL39, IFI27, ARF6, CHD11, SDPR and KRT18.
Scott Lowe (Classic Individual)
The epigenetic control of tumor ecosystem dynamics during PDAC progression
Pancreatic ductal adenocarcinoma (PDAC) is an invariably lethal cancer that arises in the context of a unique tumor microenvironment. This project aimed to examine the dynamic nature and regulation of intratumoral heterogeneity in pancreatic cancer, from its inception through metastasis. To do so, we implemented an integrative in vivo approach that combines single-cell technologies, state-of-the art computational analyses and unique mouse models of pancreatic cancer enabling perturbation of gene function at different disease stages. Our results uncovered that Kras gene mutation and tissue damage (pancreatitis) cooperatively remodel the chromatin landscape of the pancreatic epithelium to produce altered cellular states that distinguish the neoplastic process from physiological regenerative responses (Alonso-Curbelo et al., Nature 2021). Ongoing studies are incorporating novel computational methods that integrate transcriptional and chromatin information to predict cancer-initiating cells and their paths to overcoming key barriers to malignancy: initiation, benign to malignant transition, and metastatic dissemination. Collectively, our work indicates that large-scale epigenomic remodeling events integrating inputs from tissue context (inflammation) and genetics (driver gene mutations) direct the co-reprogramming of pancreatic epithelial cells and their niche early during tumor development, and establishes key cell- and non-cell-autonomous nodes that may pave the way for novel means to intercept and potentially revert the aberrant cell states responsible for PDAC initiation and metastatic progression.
Paul Chapman with Kayvan R. Keshari, Taha Merghoub, Craig B. Thompson, Santosha Vardhana, Richard White, and Jedd Wolchok (Multi-Project)
Metabolic crosstalk between melanoma and its microenvironment
There have been astounding advances in the treatment of metastatic melanoma with both targeted therapy and immunotherapy, but unfortunately more than half of people with metastatic melanoma still die of the disease. Therefore, it is important to understand the mechanisms of melanoma metastasis. This wide-ranging effort will explore the role of cellular stress and immune responses on metastasis in melanoma. Researchers also will investigate how the melanoma microenvironment induces metabolic alterations within melanoma cells that directly facilitate cellular invasion and metastasis.
This is the first GMTEC award to include multiple projects. In addition, the research team has applied to the National Institutes of Health for a P01 grant related to this work.
Project 1: Molecular mechanisms, immune responses, and cancer cell metabolism
Santosha Vardhana and Craig Thompson
This project will explore the metabolic consequences of redox regulation and its effect on T cell function within the tumor. It aims to look at the connection between metabolic adaptation in the setting of chronic inflammation and the promotion of immune evasion and metastasis in melanoma. The researchers will examine the role of altered metabolism in promoting immune dysfunction in melanoma. They also seek to determine the contribution of exogenous fatty acids to altered melanoma-specific immune responses.
Project 2: Characterize the role of lipid uptake–induced ER stress in metastasis of melanoma
Richard White
This project will examine the ability of melanoma cells to metabolize fat as an energy source despite BRAF inhibition. One tool in this research is transgenic zebrafish, which will be used to characterize the role of fat cells in the progression of melanoma. These models will also be used to assess the mechanism of lipid-mediated ER stress in melanoma invasion and study the effects that lipids have on immune cells within the tumor microenvironment. The project will also look at whether blocking SLC27A fatty acid transporter proteins has therapeutic potential, and may ultimately lead to clinical trials evaluating this approach.
Project 3: Enhancing immune-mediated control of melanoma metastasis by modulating glycolytic stress
Jedd Wolchok
This project will examine the LDH/lactate axis and the effect of lactate on immune suppression in melanoma. It aims to study the connection between lactate production and the anti-melanoma immune response. It will also investigate whether blocking lactate production can improve T cell responses against melanoma.
Mary Baylies
Identifying novel drugs that inhibit aRMS metastasis.
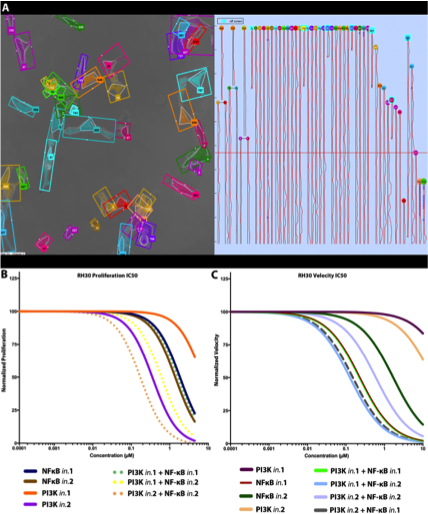
The most common pediatric soft tissue sarcoma is rhabdomyosarcoma (RMS), representing 3 to 5 percent of all childhood cancers. Alveolar RMS (aRMS) is its most aggressive form and is associated with expression of PAX3- or PAX7-FOXO1 fusion oncoproteins. Although successive clinical trials have improved survival rates for RMS patients, the outcome with standard treatment for those patients with metastatic or recurrent disease remains bleak. (The five-year survival rate is less than 20 percent.) Moreover, within the last 30 years, there have been no significant changes in treatment, and no precision treatments exist. Hence, there is a critical need to investigate RMS metastasis and to identify novel therapeutic agents for its treatment. This is the goal of the Baylies lab’s work.
With funding from MTECH, the Baylies lab has identified 17 FDA-approved or in-trial drugs that target aRMS metastasis. These drugs, combined with a novel, ongoing screen in a Drosophila aRMS model, reveal 11 pathways essential for disease development. The activity of eight of the identified drugs that affect the RAS-MAPK, PI3K-AKT, and NFkB signaling pathways has been confirmed in this in vivo model. Importantly, the Baylies lab has identified three combinations of these drugs that eliminate or significantly reduce the metastatic potential of aRMS.
Based on these and published data, the Baylies lab is testing the FDA-approved/in-trial drugs that target RAS-MAPK, PI3K-AKT, and NFkB for validation and further characterization in human RMS tumor cell lines and in an aRMS ‘metastasis’ xenograft model. For the human tumor cell lines, the Baylies lab has developed a high throughput, image-based approach in collaboration with Andrew Cohen at Drexel University to assess tumor cell behaviors upon drug treatment (single and combinatorial treatment). The Baylies lab has now identified several FDA-approved/in-trial drugs and drug combinations that preferentially affect different aspects of human metastatic tumor cell behavior, including adhesion, migration, proliferation, or survival. The Baylies lab is now examining these drug combinations in aRMS xenograft models that reproduce aspects of the metastatic process, including survival in the blood stream, extravasion, colonization of different tissue sites, and proliferation at these sites. These experiments provide the critical next step toward patient treatment. Successful completion of these experiments will provide new precision therapeutic agents and regimes for testing in the clinic.
Scott Keeney
Topoisomerase II in tumorigenesis and cytotoxic chemotherapy.
Richard White
Development of microenvironmental CRISPRs for the study of metastasis.
Johanna Joyce
Investigation of the Microenvironmental Transcriptome in Breast to Brain Metastasis.
Sarat Chandarlapaty
Role of ESR1 Mutation in Breast Cancer Metastases
Scott W. Lowe
Dissecting the interactions between Smad4 and p53 inactivation in pancreatic cancer metastasis
Kathryn Anderson
Strip1 and STRIPAK complexes in cell migration and metastasis
Mary Baylies
Finding New Treatments for Rhabdomyosarcoma
Colin Begg
Contralateral Breast Cancers: Independent Cancers or Metastases?
Richard White
Metastasis in the Zebrafish System
Robert Benezra
Neutrophil-Mediated Antimetastatic Therapy
Johanna Joyce
Developing Novel Strategies to Target Tumor Cell-Macrophage Interactions in Brain Metastasis
Daniel A. Heller
Real-Time In Vivo Marker Detection for Rapid Measurement of Metastasis Onset
Morgan Huse
Exploring the Anti-Tumor and Anti-Metastatic Function of Protein Kinase C-θ Microclusters in Natural Killer Cells
Alexander Rudensky
Regulatory T cells and Breast Cancer
David A. Scheinberg
Does Ablation of Tumor Neo-Vasculature Promote or Inhibit Metastatic Potential?
Timothy A. Chan
Large-Scale Analysis and Clinical Application of the Breast Cancer Metastasis Epigenome
Nai-Kong V. Cheung
Genetic Analyses of Secondary CNS Metastases in Neuroblastoma (NB)
Larry Norton
Identifying Clinical Tools to Detect SRC as a Critical Moderator of Bone Metastasis Latency in Women with ER Positive
Filippo Giancotti
Functional Genomics Analysis of Tumor Dormancy at Metastatic Sites
Shared Resources
-
The MSOT Acuity Echo for Clinical Optoacoustic Tomography
Moritz F. Kircher, MD, PhD -
An Image-Guided Microirradiator for Preclinical Radiation Research
Joseph Deasy, PhD -
The Ivis Spectrum for Pre-Clinical Oncology Research
Pat Zanzonico, PhD